Update February 2021. I added material relating to the DeepSig-claimed variation of the roll-off parameter in a square-root raised-cosine pulse-shaping function. It does not appear that the roll-off was actually varied as stated in Table I of [R137].
DeepSig’s datasets are popular in the machine-learning modulation-recognition community, and in that community there are many claims that the deep neural networks are vastly outperforming any expertly hand-crafted tired old conventional method you care to name (none are usually named though). So I’ve been looking under the hood at these datasets to see what the machine learners think of as high-quality inputs that lead to disruptive upending of the sclerotic mod-rec establishment. In previous posts, I’ve looked at two of the most popular DeepSig datasets from 2016 (here and here). In this post, we’ll look at one more and I will then try to get back to the CSP posts.
Let’s take a look at one more DeepSig dataset: 2018.01.OSC.0001_1024x2M.h5.tar.gz.
The dataset is from 2018 and is associated with the O’Shea paper The Literature [R137]. It, like the other two I’ve analyzed (here and here) [Update: A fourth.], is currently available on the DeepSig website. The dataset contains 24 different signals each of which is provided at each of 26 different SNR values. For each signal and SNR combination, there are 4096 instances, and each instance always has length 1024 samples. The data are stored in an HDF5 format, so I used normal HDF5 tools to extract the signals, such as h5dump. I’ll show you below how I learned all these attributes of the dataset.
When you unzip and untar the archive, you get three files: LICENSE.TXT, classes.txt, and GOLD_XYZ_OSC.0001_1024.hdf5. The latter is where the data is, and classes.txt looks like this:

The data itself is contained in the GOLD*hdf5 file, which requires a bit of examination to understand, and to attempt to connect to the classes shown in classes.txt.
I used hdf5-file tools available under linux (Ubuntu and Fedora Core in my case) to discover the structure of the data. The first step is to use h5dump to query the file about the datasets it contains. This leads to the following terse output:

So far from the description on DeepSig’s website, and the data description in the associated paper (The Literature [R137]), we know that there are 24 signals, the intended SNR range is dB, and that each individual signal data-record will always be 1024 samples in length, presumably complex-valued samples.

But we don’t know exactly how many of each signal there are nor the signal parameters, such as symbol rate. And, crucially, we don’t yet know which signal type is associated with each data record. So let’s look at the three datasets /X, /Y, and /Z.
The Datasets /X, /Y, and /Z
When we attempt to dump the contents of dataset /X, we get an output with the following header:

The DATASPACE SIMPLE line indicates that the dataset /X is three-dimensional, having 2555904 records in the first dimension, 1024 in the second, and 2 in the third. So we can guess that there are 2555904 total signals, each has 1024 samples, and those samples are complex, so need two values per sample. That explains the three dimensions.
Looking at dataset /Y, we see

So the /Y dataset is two-dimensional, with 2555904 records in the first dimension and 24 in the second. As we can see from the first few records, the 24-element vector for each record is a binary vector with only one value equal to 1 and the rest equal to 0. So this must be the modulation-type indicator. Looking good! Mysterious, and requiring some sleuthing, but good.
Turning to dataset /Z, we see

So /Z is a single vector of length 2555904 and with starting values of -20. If you look at the bottom of the vector, the value is 30, so this is the SNR-parameter dataset. There are 26 distinct SNR parameters, ranging from -20 to +30 in steps of 2. The parameter is held constant for 4096 values, then moves on to the next value. Once it gets to +30, the next value is again -20. The period of this parameter is therefore 4096*26 = 106496. To make this concrete, here are some plots of the SNR parameter:



So it looks like the SNR parameter is held constant for 4096 data records, and the SNR parameter sequence repeats after 4096*26 = 106496 data records, indicating that the signal class is likely held constant for 106496 data records. Turning back to the signal-class indicator dataset /Y, we see that the binary indicator vector switches from a one in the first location to a one in the second location exactly after 106496 data records:

The signal-class indicator dataset /Y isn’t free of mysteries, but almost. There are a few places where the indicator as output by h5dump doesn’t quite make sense:

Presumably the signal-class indicator vector position is a map into the classes.txt set of strings I showed at the top of this post. So if the vector shows a 1 in the first position, then that data record would correspond to the 32PSK signal type. How could we verify that?
Analysis of Extracted Data Records
Let’s try to analyze some of the data records we can extract from the hdf5 file and see if they have characteristics that match the corresponding signal type as determined by the associated position of the 1 in the signal-class vector of dataset /Y.
The main analysis is for a subset of data records for each signal type. I used h5dump to extract one example of each SNR condition for each signal type. Since I don’t really know the signal types in classes.txt conform to the signal-class indicator vector in the archive, I’m just going to refer to each type in terms of the offset into the hdf5 file. We know that each new signal type starts at a data-record offset of , so I’ll refer to the signals in terms of the offset
. I use h5dump commands like this:
h5dump -d /X -k 1,1,1 -s $offset,0,0 -S $stride,1,1 -c $num_blocks,1024,2 GOLD_XYZ_OSC.0001_1024.hdf5
where offset is , stride is 4096, and num_blocks is 26. I do this for all 24 offsets (starting with 0). This produces 24 data records each with length 26*1024 = 26624 complex samples.
First let’s look at the modulus of the 24 signals:

Since the SNR parameter increases to the right, but the moduli decrease (generally), the SNRs are achieved by decreasing the noise power, thereby decreasing the total power.
The closest thing to an exactly constant-modulus signal is Offset 21, but Offset 22 is also close to constant compared to the other signals. Offsets 17 and 18 are strange and non-monotonic in the modulus behavior. But not much else is evident. Let’s turn to plots of the real and imaginary parts:


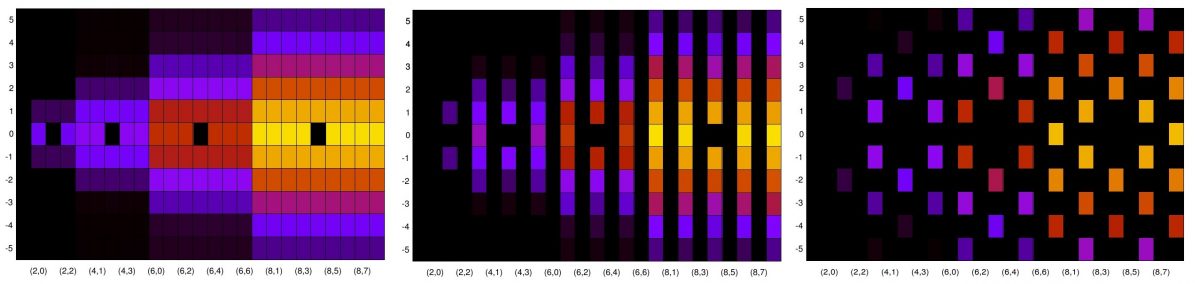
From the real and imaginary plots, we see that most of the Offsets produce approximately zero-mean sequences, but Offsets 0-2, 17, and 18 do not. Finally, let’s look at some power spectra. First I’ll show the PSDs for each signal, taking into account all 26624 samples and using the TSM for spectrum estimation:

The first three Offsets (0, 1, and 2) produce a signal with a QAM/PSK-like spectral shape and also an additive sine-wave component, like what you see for OOK signals. These are followed by 14 PSDs that look like garden-variety PSK/QAM PSDs for a signal with symbol rate of around 1/10. Those are followed by Offsets 17-21, which show signal types that are very nearly sinusoidal (Offsets 17, 18, 21) or periodic (Offsets 19 and 20). Finally, there are two more garden-variety PSK/QAM PSD shapes. Since these PSDs take into account the full range of SNRs (-20 to +30), I’ll show the PSD estimates just for the final subblock (SNR parameter of +30) to get a low-noise look at each signal:

If the mapping between the signal-class strings in classes.txt matched this data, we would expect to see an approximately constant modulus (applied channel effects can ruin the constancy of the modulus) for FM, which would be Offset 3. But the closest to constant modulus is Offset 21. Offset 22 is the next closest. So the mapping is in serious doubt.
If we return to The Literature [R137], and examine the proffered confusion matrices there, we see a different ordering of the signal classes:

This ordering is more consistent with the signal-class indicator vector in dataset /Y. The first signal is OOK, which should have a PSD with a typical QAM/PSK bump and an impulse midband, which it does in Figure 15. The final two signals in the confusion matrix are GMSK and OQPSK, which also have PSK/QAM PSDs and should not have impulses, and that is what we see in the final two PSDs (Offsets 22 and 23) in Figure 15.
Between 8ASK and AM-SSB-WC, there are 14 garden-variety types in the confusion matrix, which is consistent with Figure 15. Finally, the analog signals correspond to Offsets 17-21 in the confusion matrix and those Offsets correspond to the PSDs in Figure 15 that are the most non-PSK/QAM in appearance.
So the mapping provided by DeepSig in classes.txt is incorrect, but a correct one is possibly

To check that the rather severe subsampling of the data records for each modulation offset above didn’t miss anything significant, I extracted every tenth 1024-point subblock from the archive for Offset = 3. Here are the PSDs:
Square-Root Raised-Cosine Roll-Offs in the Dataset
In [R137] we find the table of random variables that are applied to the signals in Table I:

In Table I, is the variable the represents the roll-off parameter (don’t confuse it with cycle frequency!), so that the dataset should exhibit signals with excess bandwidths in the range of
%. To check this I would like to pass examples through a modulation-recognizer that is sensitive to pulse-shaping functions, but the authors’ preoccupation with very short data records inhibits this approach. Let’s see how far we can get by looking at some PSDs.
I estimated the PSDs for the strongest Offset-3 signals and plotted them on a single set of axes as shown in Figure 19. Overlaid on those
PSDs are three additional PSDs corresponding to PSK signals with roll-offs
of
and
. (All estimates use identical methods and spectral resolution parameters.) I have to conclude that either the range of roll-offs is much smaller than claimed or the pulse-shaping function used by DeepSig is not the usual square-root raised-cosine pulse function. To plot the three colored PSD lines in Figure 19 requires that I chose a symbol rate. My best guess is exactly
because when I pass the
-point blocks through a blind non-conjugate cycle-frequency estimator (such as the FAM or SSCA), the only consistently detected value is
.

Discussion
The impulses in 4ASK and 8ASK (Offsets 1 and 2) are expected because the constellations were probably all non-negative numbers along the real-axis, which is a typical definition of ASK, so that checks.
The five analog waveforms don’t make sense to me. They are essentially impulsive in the frequency domain, and there doesn’t appear to be much difference between those “with carrier” and those with “suppressed carrier.” In particular, suppressed-carrier signals should not contain impulses in their spectra. It looks like the AM and FM signals are being driven by a sinusoidal message signal (non-random).
Otherwise, this is a much better dataset than the other two DeepSig datasets I’ve analyzed (see All BPSK Signals and More on DeepSig’s RML Datasets).
It still suffers from the “one BPSK signal” flaw, because it looks like the symbol rate and carrier offset never significantly change (see Table I in The Literature [R137]), and I have come to the conclusion that the excess bandwidth (roll-off parameter for the square-root raised-cosine pulse-shaping filter) is not actually varied. (Compare this to my Challenge Dataset.)
The dataset also suffers from the preoccupation with very short data records. This prevents verification and other kinds of analysis and comparison. If the data records were made longer, presumably a machine learner could still train and test using a subset of each data record (use the first samples of each data record with length
), so there isn’t any disadvantage in making longer records except the size of the archive increases.
As usual, let me know if I’ve erred or if you have a relevant comment in the Comments section below.
Hello Chad,
thanks for that interesting analyses of the radio ML datasets. Assuming these are correct: This would be a serious issue to the scientific community working in that topic! Have you received any statement or comment from Tim O’Shea?
Thanks for stopping by and leaving a comment Peter! Welcome.
The first post I did on Machine Learning for Modulation Recognition was a critical review of the RML paper by O’Shea, Clancy, and Corgan. That was in early 2017. Tim left this comment, and you can read my reply to that comment, which was never answered. Subsequently, I met Tim and tried to work on some ML stuff with him. I know for a fact he has my Data Set for the Machine Learner’s Challenge. But he never disclosed to me his results.
Since then, he hasn’t commented on any of the data-set analysis posts here, here, and here. This is all fine with me.
Hi Mr. Spooner. I am Electrical&Electronics Engineering 4th grade student in Ankara University. My project homework is about over the air deep learning based radio signal classification. And i see that you have worked on this paper and written its codes. Could you please share the python codes? If you answer me positively, then i will be thankful to you.
Fatih: I’m sorry to tell you that you are mistaken. I did not work on the paper associated with this data set ([R137] in The Literature). I did analyze the data set for the benefit of the modulation-recognition community. I suppose you could try contacting the authors of [R137].
I see, thank you sir.
Hello Mr. Spooner!
I am currently pursuing my Phd. on the topic of Spectrum Sensing and I found valuable information in your work. Your analysis on the DeepSig dataset is the most detailed I’ve found so far. I have noticed that in this article you state that each modulation class is maintained for 106496 samples in the /Y dataset ( also the SNR is ranged from -20 to +30 in the /Z dataset for 106496 samples ) , but each signal you extracted from the /X dataset has only 26624 samples. Are there multiple instances of the same signal for each SNR (4096 instances), from which you extracted only one instance or am I missing something? Please clarify this aspect for me. Also I am looking forward to tackle the dataset you provided in the “Data Set for the Machine-Learning Challenge” article. Thank you!
Welcome to the CSP Blog Mutescu (or is it Marian?)!
I will answer your question. But first, I need you to answer one of mine. You say
I have seen zero other analyses of the DeepSig data sets. I’ve seen only applications of ML to the data set.
Can you please write down here links to or citations to the other analyses that you’ve seen?
You are right, there are zero official analyses on this dataset. The other “analyses” I mentioned were a couple of answers and opinions scrambled trought forums.
I think you’re asking about Figure 12. I extract one example of each modulation type (what I’m calling the offset in the post) for each SNR value. I then concatentate those data records. So for a particular signal type (offset), I have 26 (there are 26 SNRs) complex vectors with length 1024. Concatenating these 26 vectors leads to a complex vector with length 26*1024 = 26624. I then plot the magnitude (modulus) of that vector on one of the axes in Figure 12. So each little subplot in Figure 12 shows 26 different versions of a single signal type, just with different SNRs.
Yes, there are multiple instances of the same signal for each SNR, but I only extracted one instance of each SNR for each signal to create the 24 subplots in Figure 12.
For the video of power spectrum estimates near the end of the post, I extract lots of instances of each SNR but for only one signal type (offset 3).
Does that clarify the post for you? Let me know…
Yes, thank you for taking your time to answer my questions!
Hi Dr. Spooner!
The captions for Fig. 13 and 14 are virtually the same. Is Fig. 14 the imaginary part?
Welcome to the CSP Blog Laura! Thanks for the comment.
Yes, Figure 14 is the imaginary part, so I fixed the caption to reflect that. I think I did a copy-and-paste error. Ouch.
Let me know if you find other errors!
Hello, I have some questions to ask about the DeepSig dataset
1. In 2018.01A, the signal is represented as 2 * 1024, while in 2016.10a it is 2 * 128. Is the second dimension, which is the time series in the article, downsampling from 1024 to 128? I wonder if it is a set of data directly sampled, or different collection points, or different collection samples?
2. In the introduction paper of the original author’s dataset, channel conditions were mentioned. So, when dividing the training and testing sets, was there a division based on channel conditions, and was this reflected in the dataset?
Welcome, bravelykiwi, to the CSP Blog! Thanks for the comment.
I have also analyzed the other DeepSig RML datasets (2016a, 2016b, 2016c). You can probably answer your questions by studying all of these posts. (Also, for all of my posts on the RML datasets, including some commentary, go to the right side of any CSP Blog page, find the Categories list, and click on RML.)
I doubt that the 128-sample signals in the 2016 datasets arise from downsampling the 1024-sample signals in the 2018 dataset. Note that the signals in all RML datasets have occupied bandwidths of about 0.17 (normalized) Hz. If you try to downsample the 2018 signals by 8 (because 128 = 1024/8), you’d violate the sampling theorem and you’d end up with distorted signals. Perhaps more practically speaking, the 2018 signals were posted two years after the 2016 signals. It is exceedingly unlikely that the 2018 signals were created in 2016, the 2016 signals were constructed from those signals, and then the original signals were eventually posted two years later as the 2018 dataset.
But, overall, this is a question for the machine learners over at DeepSig.
As I mention in the post (and in the other RML posts), there doesn’t seem to have been any significant application of channel effects to any posted DeepSig signals. (For example, see Figure 19 in the post.) So I doubt there is any significant dataset organization or division on the basis of channel effects.
Again, though, this might be a question that is better directed at DeepSig, since it asks about intent.
Hi Chad,
What I got from the last paragraph of this post (please correct me if I am wrong), the signal length of 1024 samples is not long enough to (reliably) estimate cyclostationary statistics such as spectral correlation function, and other higher-order cyclic cumulants? If so, how to know how large the signal should be to get reliable estimates? Thanks.
Hey Ishtiaq! Thanks for reading the CSP Blog and the comment!
Start here: The resolution product.
Hi,
I am currently conducting some research on modulation recognition using the RML2018.01A dataset and now I need its sampling rate. I didn’t find it in the thesis. Could you please tell me? This is very important to me. I’m very sorry to disturb you.
best regards
Yu Sun
Welcome to the CSP Blog Yu Sun! Thanks for the comment.
Looking back at my analysis of this dataset, I confirmed my remembrance of a unity sampling rate. That is, this dataset does not come with a physical sampling rate as far as I can tell. If it was created with the aid of SDRs and gnuradio, then there would be a physical sampling rate associated with that. But once the signals are sampled and downconverted to complex baseband, the data can be described in terms of normalized times and frequencies.
So I think the answer is “1.0 Hz.”
Hi,
I divided the 2018.01A dataset into 6,2,2 and trained a model, which performed well in the test set. Now I have collected a batch of independent test sets myself and want to test the effect with the previously trained model. It was found that the generalization effect was very poor, and basically all predictions were incorrect. I don’t understand where the problem lies. Could it be caused by different sampling rates or else? Could you also tell me about the sampling rate of 2018.01a? I’m very sorry to disturb you.
best regards
You’ll need to explain this in detail.
Indeed, the RML 2018 dataset appears to contain digital signals with a single symbol rate (1/8) and analog signals that do not conform to standard models associated with their labels (AM-DSB, FM, etc.).
But there are many random variables involved in creating an RF-signal dataset, and there could be differences in their probability density functions between your dataset and RML 2018. So you’ll have to look carefully at your data, just as we all need to look carefully at publicly available datasets.
Have you seen my sequence of papers (and CSP-Blog posts) on ML-based modulation recognition and the pursuit of high generalization?
[50] J. A. Snoap, D. C. Popescu, and C. M. Spooner, “On Deep Learning Classification of Digitally Modulated Signals Using Raw I/Q Data,” IEEE Consumer Communications and Networking Conference, Jan. 2022.
[51] J. A. Latshaw, D. C. Popescu, J. A. Snoap, and C. M. Spooner, “Using Capsule Networks to Classify Digitally Modulated Signals with Raw I/Q Data,” The 14th International Conference on Communications (COMM2022),” Bucharest, Romania, June 2022.
[52] J. A. Snoap, J. A. Latshaw, D. C. Popescu, and C. M. Spooner, “Robust Classification of Digitally Modulated Signals Using Capsule Networks and Cyclic Cumulant Features,” Proceedings of the Conference on Military Communications (MILCOM), Rockville, MD, November 2022.
[54] J. A. Snoap, D. C. Popescu, J. A. Latshaw, and C. M. Spooner, “Deep-Learning-Based Classification of Digitally Modulated Signals Using Capsule Networks and Cyclic Cumulants,” MDPI Sensors, June 2023, https://www.mdpi.com/1424-8220/23/12/5735/pdf, DOI: 10.3390/s23125735.
[55] J. A. Snoap, D. C. Popescu, and C. M. Spooner, “Novel Nonlinear Neural-Network Layers for High Performance and Generalization in Modulation-Recognition Applications,” Proceedings of the Conference on Military Communications (MILCOM), Boston, MA, November 2023.
[56] J. A. Snoap, D. C. Popescu, and C. M. Spooner, “Deep-Learning-Based Classifier with Custom Feature-Extraction Layers for Digitally Modulated Signals,” IEEE Transactions on Broadcasting, Vol. 70, No. 3, pp. 763-773, Sept. 2024.
https://cyclostationary.blog/2024/07/17/final-snoap-doctoral-work-journal-paper-my-papers-56-on-novel-network-layers-for-modulation-recognition-that-generalizes/
https://cyclostationary.blog/2023/08/15/the-next-logical-step-in-cspml-for-modulation-recognition-snoaps-milcom-23-paper-preview/
https://cyclostationary.blog/2023/06/20/latest-paper-on-csp-and-deep-learning-for-modulation-recognition-an-extended-version-of-my-papers-52/
https://cyclostationary.blog/2022/11/02/neural-networks-for-modulation-recognition-iq-input-networks-do-not-generalize-but-cyclic-cumulant-input-networks-generalize-very-well/
https://cyclostationary.blog/2022/05/18/some-concrete-results-on-generalization-in-modulation-recognition-using-machine-learning/
Dear Chad,
thank you very much for taking the time to answer my question. I’m currently in my first year of postgraduate studies, so I’m a bit ignorant about some issues. Please excuse me. Here’s the thing: One of my supervisor’s projects requires training a modulation recognition model, and then they can directly recognize the signals received in reality. To achieve this goal, I trained the model at RML2018.01a. The project manager provided me with a batch of signal data (the independent test set mentioned above), which only contained modulation labels and no other information. I have read your article these two days and I find it very meaningful than any paper on AMR I have read before (they seem to only focus on accuracy). In the article “Deep-Learning-Based Classification of Digitally Modulated Signals Using Capsule Networks and Cyclic Cumulants” I noticed that you provided a blind estimation process for CC features. If I want to successfully achieve the effect of this blind estimation process and the generalization ability of the model in this paper, what parameter information of this batch of signals should I at least need? At the same time, should I ensure that the probability density functions followed by the parameters of the two datasets, except for carrier offset and symbol rate, are as similar as possible? If they are not the same, will the generalization effect vary a lot? How do you think I should fulfill the requirements of this project? Thank you again for your reply. I’m extremely grateful.
best regards
YuSun
I question whether this is a good idea or not. What do you think of the analog signals in the RML 2018 dataset? Do you agree with my analysis that there is a single symbol rate represented in the dataset? And a single square-root raised-cosine filter?
What procedure did you or anyone else use to determine whether these signals and associated labels are correct?
You need to develop a full understanding of the signals that you intend to classify (“domain expertise“) and you need to develop a full understanding of signal processing and cyclostationary signal processing (“domain expertise”). After that, you need to fully understand the neural network model you want to use, and all of its available hyperparameters. The first two of these three things can be obtained by careful study of the CSP Blog posts, starting from the first one and reading/studying in chronological order (see here).
That’s entirely up to you and your research group. To understand which random variables are present, and which are relatively important, you will have to develop a model of radio-signal reception. What happens from the transmitter all the way to the I/Q samples? How will you select filters (ideal and/or practical)? How will you select the sampling rate of the complex envelope which determines the samples per symbol? How will you convert to complex baseband, and what will the likely range of carrier frequency offsets be? Etc.
That’s the question you’ll be trying to answer.
Contrary to the machine-learner’s mantras, there is no escaping domain expertise. At an absolute minimum, you need to know that your labels match the reality of the signals.
Dear Chad, I feel this is not a good idea either, but I haven’t thought of any other solutions yet. I agree with your analysis. I attempted to draw the spectrograms, constellation diagrams, envelope diagrams, and phase change diagrams of these signals to try to determine the correctness of these labels (is this correct?)” . If time permits, I also want to try to have a comprehensive understanding of signal processing and loop stationarity processing, but I don’t have much time left. My financial situation and my supervisor’s urging forced me not to stop and study step by step. This is a common problem in China. Thank you sincerely for your advice to a stranger. The project’s owner did not provide me with the process of generating these signals. They only asked me to train a model capable of accurately identifying signals in reality. They won’t care how difficult this will be.
My supervisor asked me to make another attempt (I couldn’t stop it), asking me to use VSG60A and RSA306B to collect some signals to expand RML2018.01A and obtain a more comprehensive dataset (although I think these two datasets cannot be merged due to the influence of parameters). Then conduct training to try to obtain better test results. Dear Chad, what can help me?
Yes, those are definitely good things to do, especially the constellation diagrams (plots) if the constellation is extracted from the signal after it is generated, rather than before.
I’m sorry to have to say that this is a recipe for failure.
Using a Signal Hound signal generator (for creating signals) and a real-time spectrum analyzer (for partial validation of each signal) is definitely a good step. Be careful with all signal generators (hardware-based and software-based) because there are pitfalls, such as the use of a modulating information sequence that is very short and repeated over and over, which does not model the real-world situation of data transmission.
There are no shortcuts! This is a complicated and complex problem area, and pretending that it isn’t won’t help your sponsor.
Dear Chad,
I’m sorry to disturb you again. I’d like to ask you some questions about generating AMR datasets. I want to use VSG60A and RSA306B to generate an AMR dataset for everyone to use. I noticed that the software provided by VSG60A contains the functions of adding carrier offset and root site-cosine as well as SNR. Should I use these parameters in the software? Should I write code to add these parameters after generating “clean” signals. If I want to use this generated dataset to publish in EI conferences (is this difficult?)”. My current idea is that it has the following 21 modulation types, specifically including [‘1024QAM’, ’16FSK’, ’16PSK’, ’16QAM’, ‘256QAM’, ‘2ASK’, ‘2FSK’, ‘4FSK’,’64QAM’, ‘8FSK’, ‘8PSK’, ‘AM’ and ‘BPSK’, ‘D8PSK’, ‘DBPSK’, ‘DQPSK’, ‘FM’, ‘OFDM’, ‘OQPSK’, ‘PI4DQPSK’, ‘QPSK’]. Among them, each modulation method has 4096 samples, each sample has 1024 sampling points or more, with a signal-to-noise ratio ranging from -20 to +30dB, and a step size of 2dB. I have no experience in the selection of carrier offset and root raised-cosine and Rs, as well as many other parameters. What do you think these parameters should be like? What scale or specific requirements should such published datasets have?
Best Regards
Yu Sun
This isn’t about CSP, or comparing alternatives like machine learning to CSP, so it is starting to be outside of my remit.
I think you and your research group need to decide what research problem you want to study, and then decide how you want to do that.
If you then run into problems with CSP, SP, or the comparison of ML results to CSP/SP results, I can usefully comment at that time.